Journal of Peking University (Health Sciences) ›› 2026, Vol. 58 ›› Issue (2): 290-300. doi: 10.19723/j.issn.1671-167X.2026.02.011
Previous Articles Next Articles
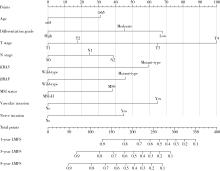
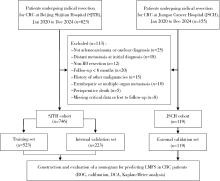
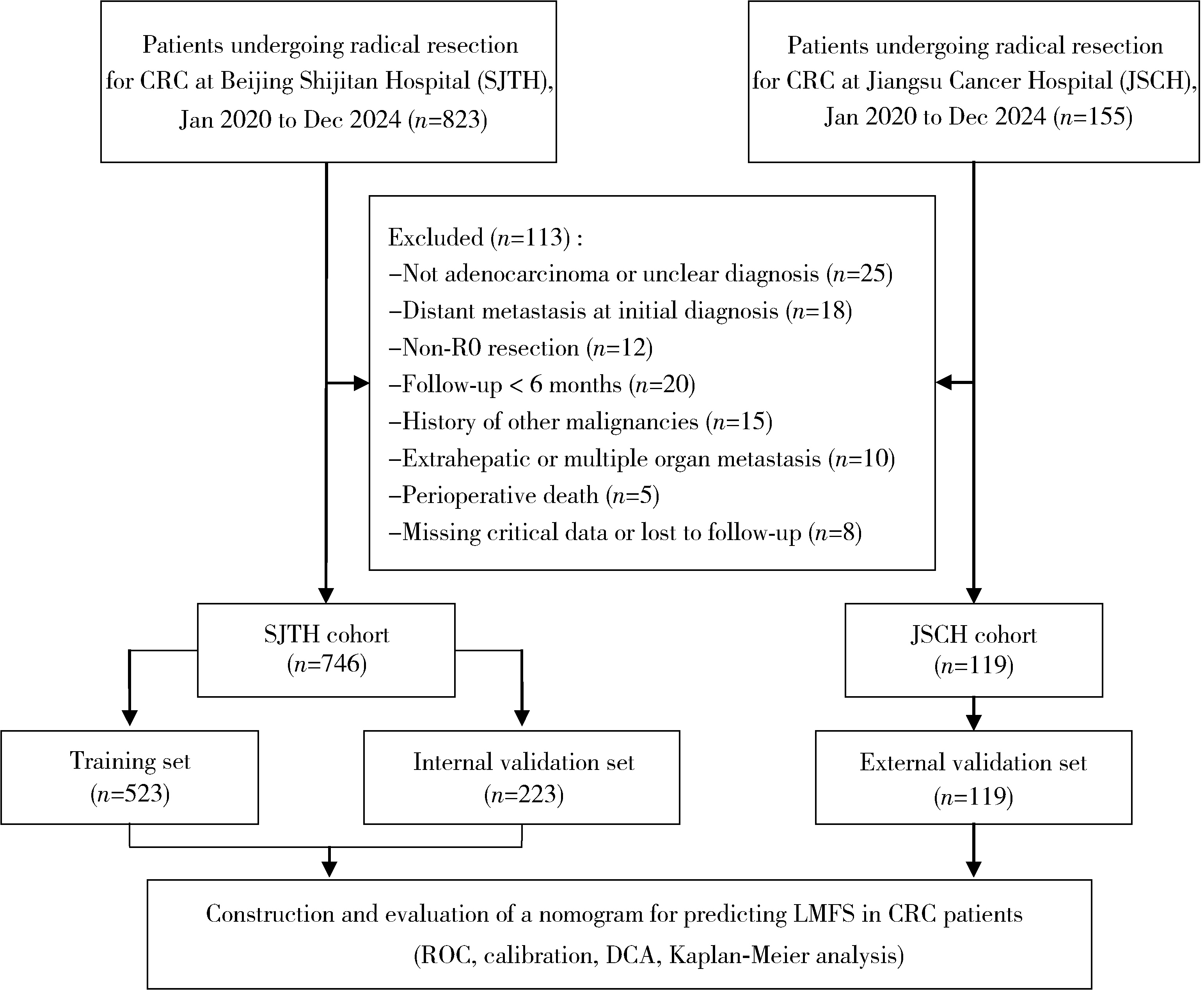
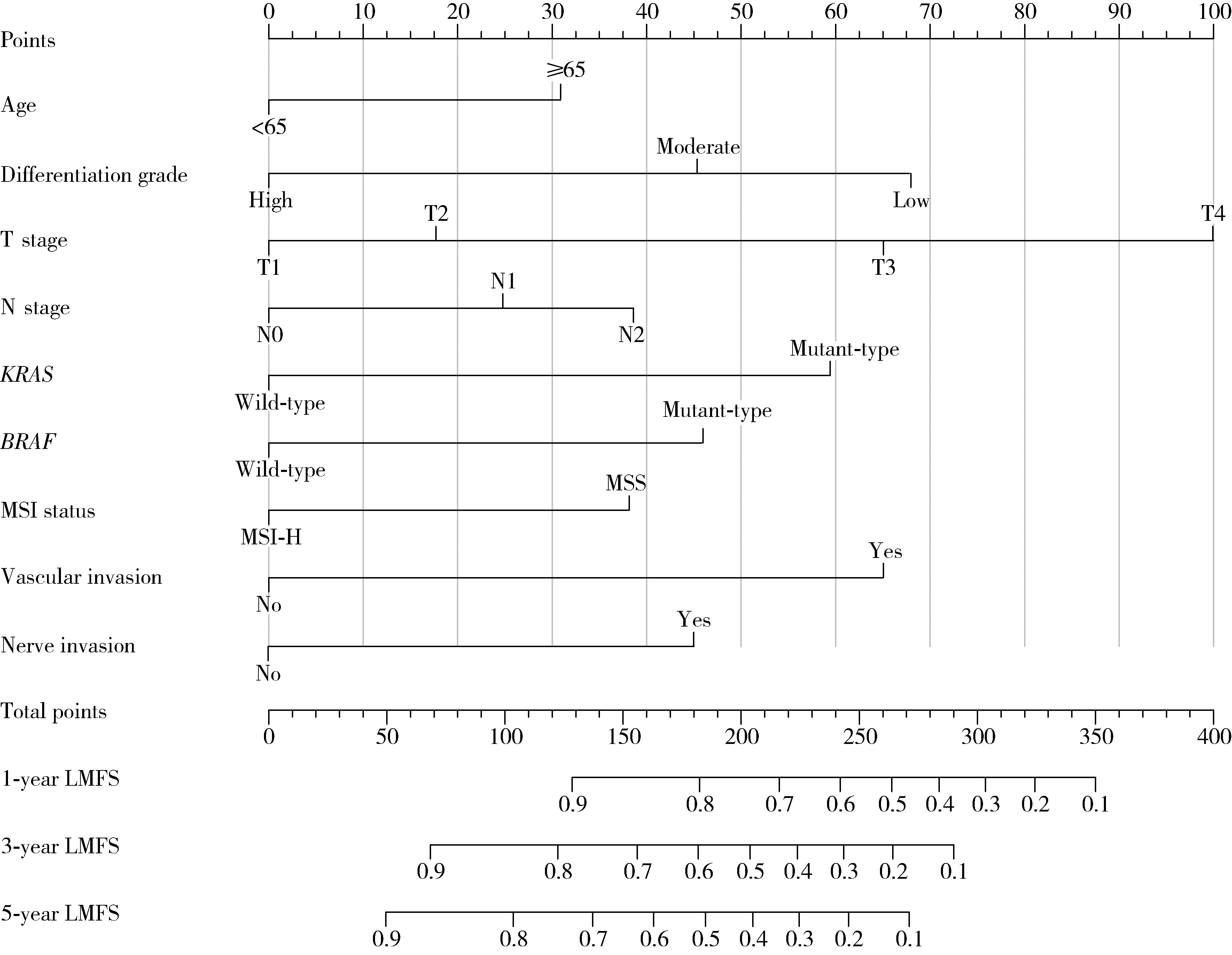
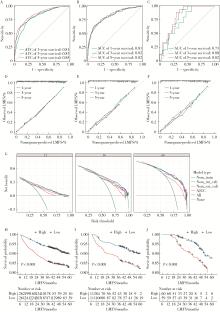
Development and validation of a multicenter nomogram predicting the risk of liver metastasis after curative resection of colorectal cancer
Nannan WANG1,2, Dajin YUAN1, Yubing ZHU1, Lei DING1,*( )
)
- 1. Department of Gastrointestinal Surgery, Beijing Shijitan Hospital, Capital Medical University, Beijing 100038, China
2. Department of Colorectal Surgery, The Affiliated Cancer Hospital of Nanjing Medical University & Jiangsu Cancer Hospital & Jiangsu Institute of Cancer Research, Nanjing 210009, China
CLC Number:
- R735.34
| 1 |
|
| 2 |
|
| 3 |
doi: 10.1200/JCO.22.01690 |
| 4 |
doi: 10.1001/jamanetworkopen.2022.36666 |
| 5 |
doi: 10.1016/j.critrevonc.2023.103990 |
| 6 |
doi: 10.6004/jnccn.2021.0012 |
| 7 |
|
| 8 |
doi: 10.1016/j.surg.2022.10.014 |
| 9 |
doi: 10.1016/S1470-2045(14)71116-7 |
| 10 |
doi: 10.1186/s12967-019-1940-1 |
| 11 |
doi: 10.1097/JS9.0000000000000415 |
| 12 |
doi: 10.7150/jca.46155 |
| 13 |
|
| 14 |
doi: 10.1186/s12957-022-02558-6 |
| 15 |
doi: 10.3389/fonc.2022.844067 |
| 16 |
Shao S, Tian D, Li M, et al. Survival prediction in sigmoid-colon-cancer patients with liver metastasis: A prospective cohort study[J/OL]. JNCI Cancer Spectr, 2024, 8(5): pkae080[2025-11-01]. https://doi.org/10.1093/jncics/pkae080.
|
| 17 |
|
| 18 |
doi: 10.21037/jgo-2025-415 |
| 19 |
doi: 10.1038/s41467-018-04278-6 |
| 20 |
doi: 10.1186/s13045-024-01627-5 |
| 21 |
doi: 10.1001/jamasurg.2018.0996 |
| 22 |
doi: 10.1038/s41408-023-00833-7 |
| 23 |
doi: 10.1093/annonc/mdz139 |
| 24 |
doi: 10.1016/j.tranon.2024.101997 |
| 25 |
|
| 26 |
doi: 10.1016/j.ijsu.2017.04.068 |
| [1] | Bowen LI, Qiang ZHANG, Yixin SUN. Establishment and validation of a risk prediction model for scoliosis after Nuss procedure in children and young adults with pectus excavatum [J]. Journal of Peking University (Health Sciences), 2025, 57(5): 941-946. |
| [2] | Zhicun LI, Tianyu WU, Lei LIANG, Yu FAN, Yisen MENG, Qian ZHANG. Risk factors analysis and nomogram model construction of postoperative pathological upgrade of prostate cancer patients with single core positive biopsy [J]. Journal of Peking University (Health Sciences), 2024, 56(5): 896-901. |
| [3] | Yuanmei LIU, Yicheng FU, Jingxin HAO, Fuchun ZHANG, Huilin LIU. Construction and validation of a nomogram for predicting in-hospital postoperative heart failure in elderly patients with hip fracture [J]. Journal of Peking University (Health Sciences), 2024, 56(5): 874-883. |
| [4] | Zezhen ZHOU,Shaohui DENG,Ye YAN,Fan ZHANG,Yichang HAO,Liyuan GE,Hongxian ZHANG,Guoliang WANG,Shudong ZHANG. Predicting the 3-year tumor-specific survival in patients with T3a non-metastatic renal cell carcinoma [J]. Journal of Peking University (Health Sciences), 2024, 56(4): 673-679. |
| [5] | Hailong HE,Qing LI,Tao XU,Xiaowei ZHANG. Construction of a predictive model for postoperative pain relief after microscopic spermatic cord surgery for spermatic cord pain [J]. Journal of Peking University (Health Sciences), 2024, 56(4): 646-655. |
| [6] | Junqi SU,Xiaoying WANG,Zhiqiang SUN. Establishment and verification of a prognostic nomogram for survival of tongue squamous cell carcinoma patients who underwent cervical dissection [J]. Journal of Peking University (Health Sciences), 2024, 56(1): 120-130. |
| [7] | Hai MAO,Fan ZHANG,Zhan-yi ZHANG,Ye YAN,Yi-chang HAO,Yi HUANG,Lu-lin MA,Hong-ling CHU,Shu-dong ZHANG. Predictive model of early urinary continence recovery based on prostate gland MRI parameters after laparoscopic radical prostatectomy [J]. Journal of Peking University (Health Sciences), 2023, 55(5): 818-824. |
| [8] | DING Ting-ting,ZENG Chu-xiong,HU Li-na,YU Ming-hua. Establishment of a prediction model for colorectal cancer immune cell infiltration based on the cancer genome atlas (TCGA) database [J]. Journal of Peking University (Health Sciences), 2022, 54(2): 203-208. |
| [9] | MU Dong-liang,XUE Cheng,AN Bin,WANG Dong-xin. Epidural block associated with improved long-term survival after surgery for colorectal cancer: A retrospective cohort study with propensity score matching [J]. Journal of Peking University (Health Sciences), 2021, 53(6): 1152-1158. |
| [10] | Wen-peng WANG,Jie-fu WANG,Jun HU,Jun-feng WANG,Jia LIU,Da-lu KONG,Jian LI. Clinicopathological features and prognosis of colorectal stromal tumor [J]. Journal of Peking University (Health Sciences), 2020, 52(2): 353-361. |
| [11] | Xue-hua ZHU,Ming-yu YANG,Hai-zhui XIA,Wei HE,Zhi-ying ZHANG,Yu-qing LIU,Chun-lei XIAO,Lu-lin MA,Jian LU. Application of machine learning models in predicting early stone-free rate after flexible ureteroscopic lithotripsy for renal stones [J]. Journal of Peking University(Health Sciences), 2019, 51(4): 653-659. |
| [12] | Le GAO,Shu-qing YU,Ji-chun YANG,Jun-ling MA,Si-yan ZHAN,Feng SUN. Quality assessment of global guidelines on colorectal cancer screening [J]. Journal of Peking University(Health Sciences), 2019, 51(3): 548-555. |
| [13] | LIU Yu-qing, LU Jian, HAO Yi-chang, XIAO Chun-lei, MA Lu-lin. Predicting model based on risk factors for urosepsis after percutaneous nephrolithotomy [J]. Journal of Peking University(Health Sciences), 2018, 50(3): 507-513. |
|
||