Journal of Peking University (Health Sciences) ›› 2026, Vol. 58 ›› Issue (2): 393-398. doi: 10.19723/j.issn.1671-167X.2026.02.026
Previous Articles Next Articles
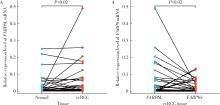
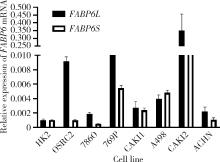
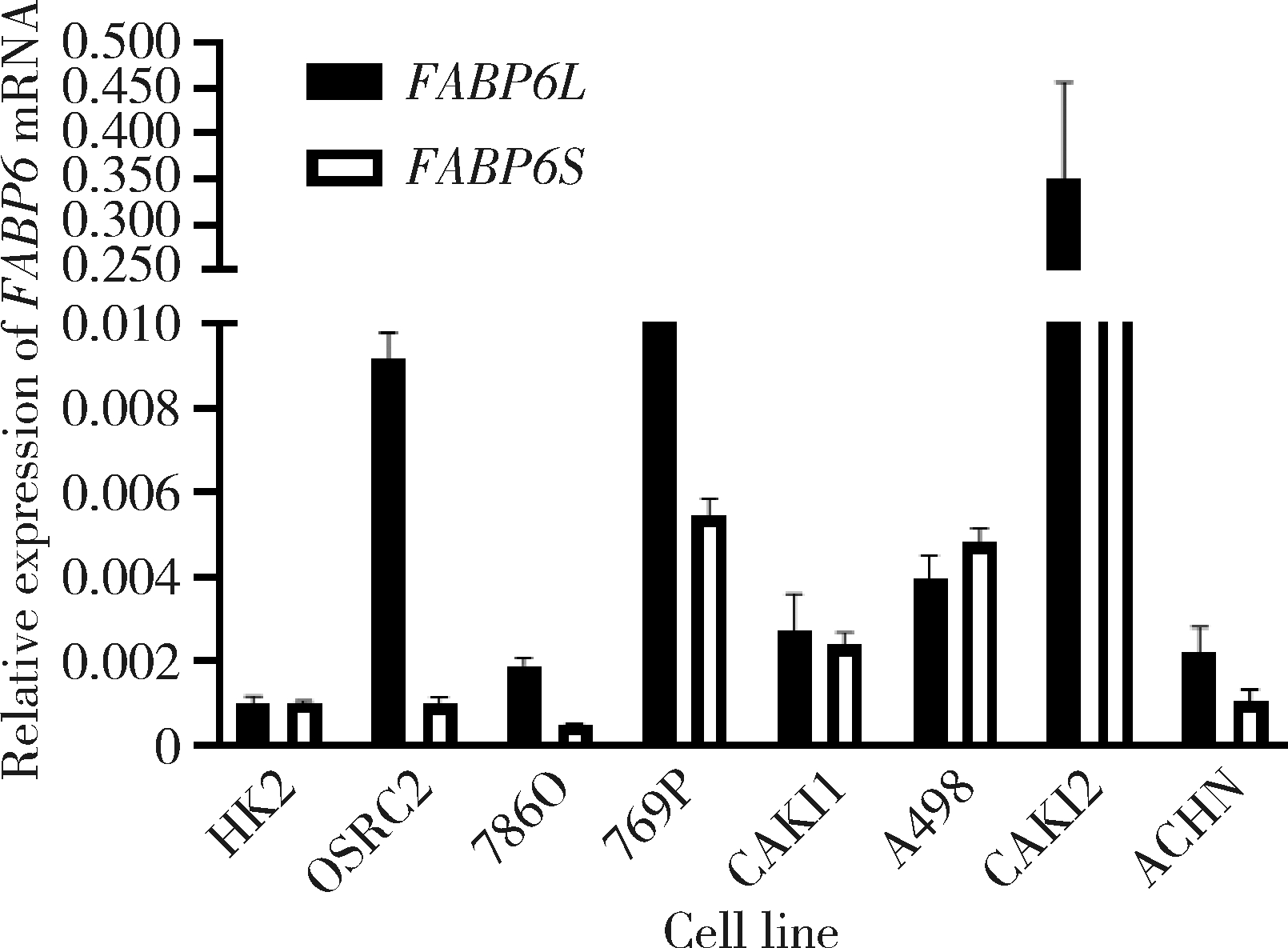
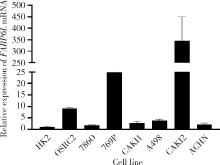
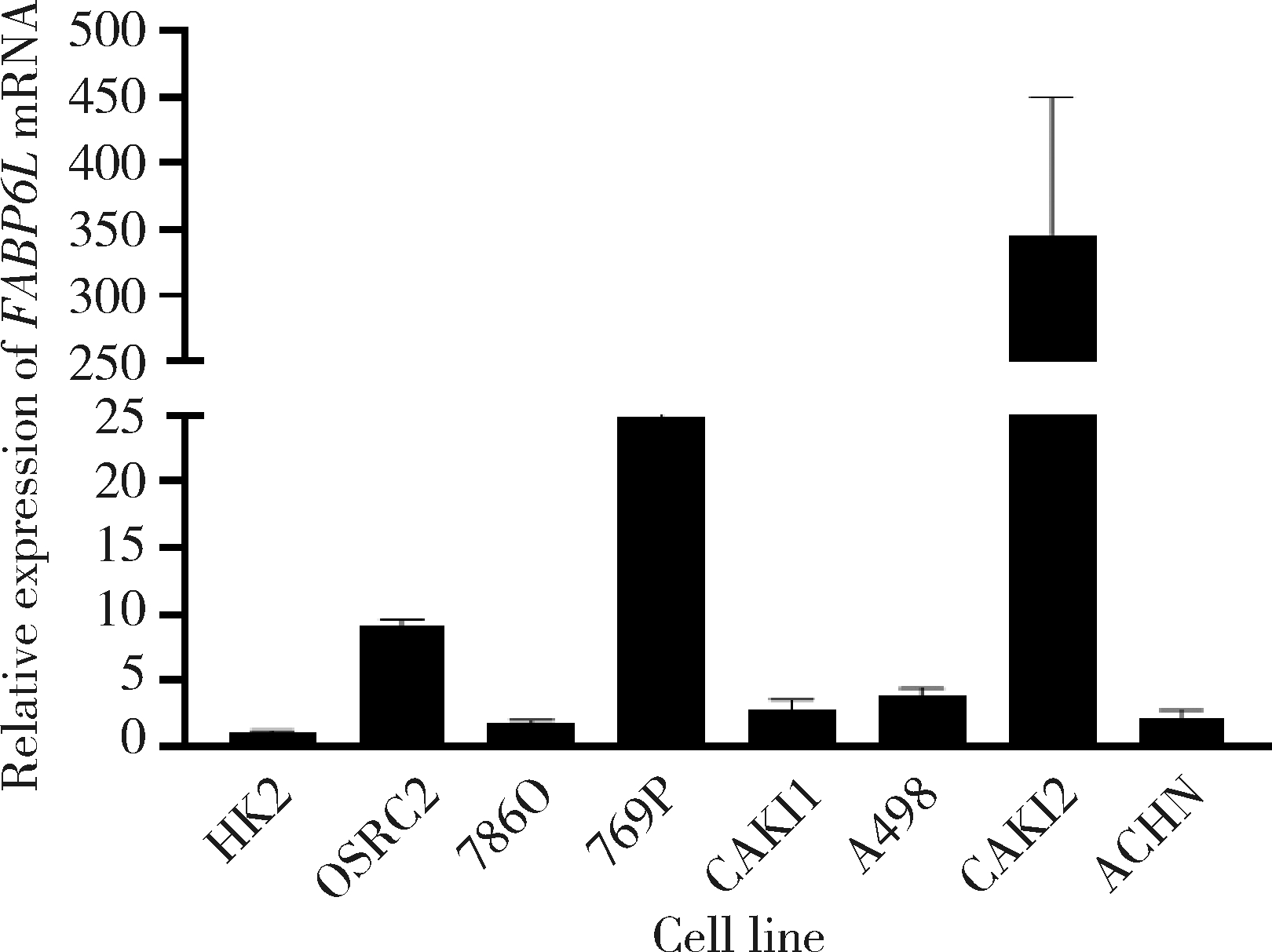
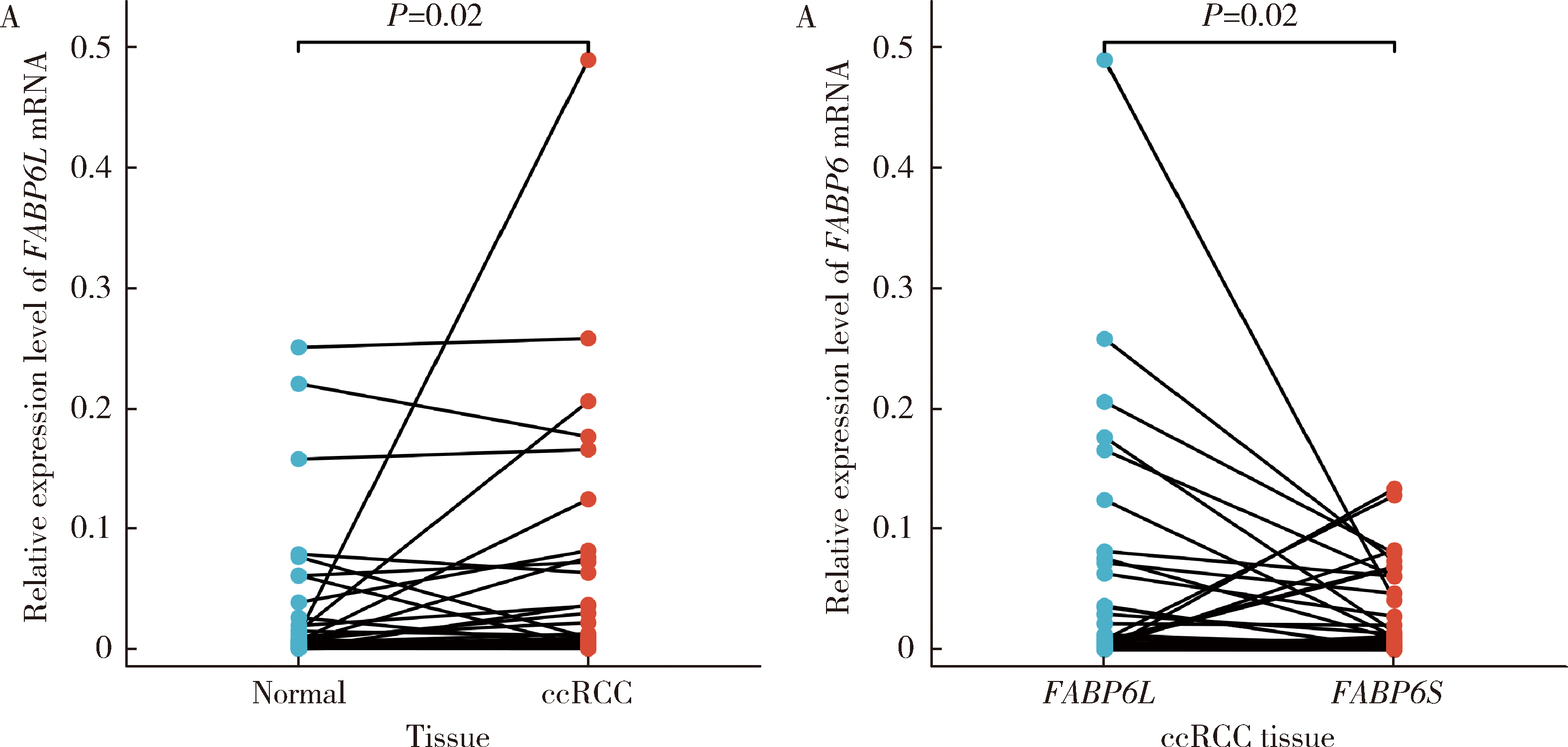
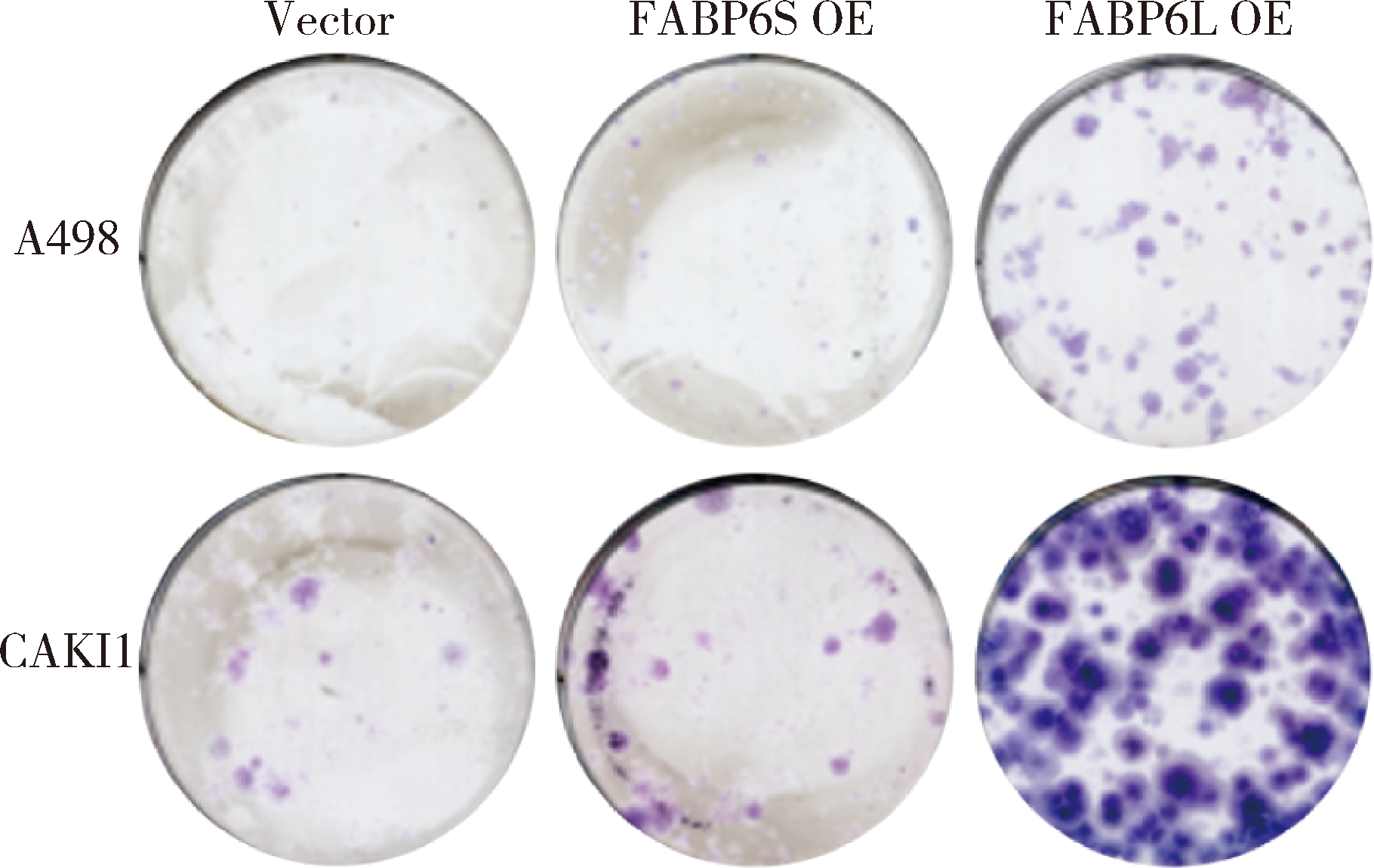
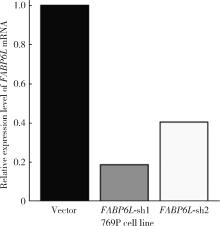
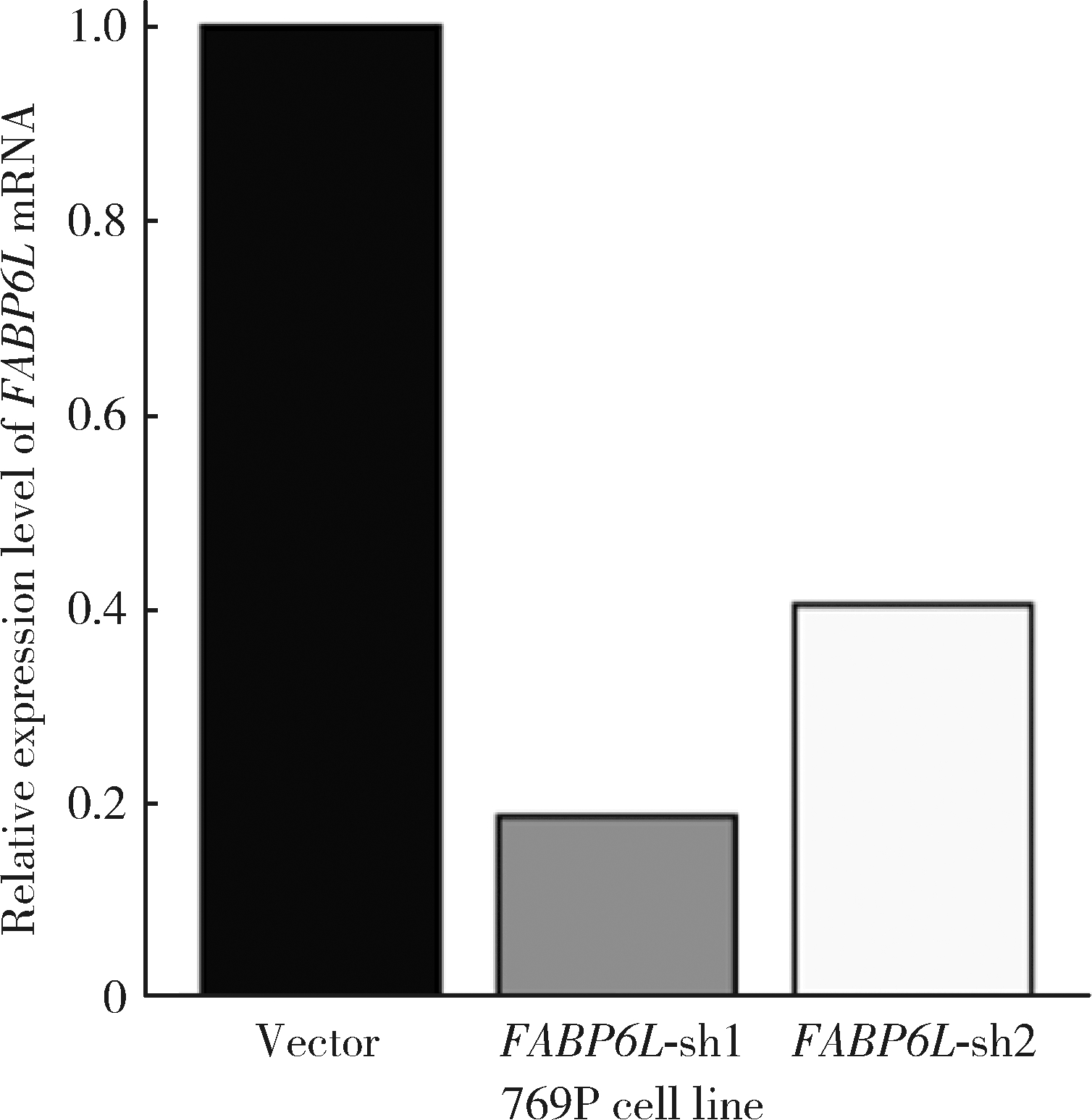
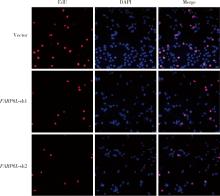
Expression and significance of the FABP6 long transcript in clear cell renal cell carcinoma
Haoming YIN1, Zijie WANG1, Fan SHU1, Zhanyi ZHANG1, Hui LIANG2,*( ), Shudong ZHANG1,*(
), Shudong ZHANG1,*( )
)
- 1. Department of Urology, Peking University Third Hospital, Beijing 100191, China
2. Department of Pathology, Peking University School of Basic Medical Sciences, Beijing 100191, China
CLC Number:
- R737.1
| 1 |
doi: 10.1038/s41588-024-01662-5 |
| 2 |
doi: 10.1158/0008-5472.CAN-06-3690 |
| 3 |
doi: 10.1016/j.bbagen.2017.07.004 |
| 4 |
|
| 5 |
doi: 10.1186/s12957-019-1714-5 |
| 6 |
doi: 10.18632/aging.205448 |
| 7 |
doi: 10.3390/cells10102782 |
| 8 |
doi: 10.1016/j.prostaglandins.2023.106732 |
| [1] | Shaohai TANG, Baoming YANG, Jiankun LI, Lili ZHAO, Yifan WANG, Shunxiang WANG. HDAC2-mediated H3K27 acetylation promotes the proliferation and migration of hepatocellular carcinoma cells [J]. Journal of Peking University (Health Sciences), 2025, 57(5): 884-894. |
| [2] | Yao ZHANG,Jinxin GUO,Shijia ZHAN,Enyu HONG,Hui YANG,Anna JIA,Yan CHANG,Yongli GUO,Xuan ZHANG. Role and mechanism of cysteine and glycine-rich protein 2 in the malignant progression of neuroblastoma [J]. Journal of Peking University (Health Sciences), 2024, 56(3): 495-504. |
| [3] | Yun-chong LIU,Zong-long WU,Li-yuan GE,Tan DU,Ya-qian WU,Yi-meng SONG,Cheng LIU,Lu-lin MA. Mechanism of nuclear protein 1 in the resistance to axitinib in clear cell renal cell carcinoma [J]. Journal of Peking University (Health Sciences), 2023, 55(5): 781-792. |
| [4] | Tian-yu CAI,Zhen-peng ZHU,Chun-ru XU,Xing JI,Tong-de LV,Zhen-ke GUO,Jian LIN. Expression and significance of fibroblast growth factor receptor 2 in clear cell renal cell carcinoma [J]. Journal of Peking University (Health Sciences), 2022, 54(4): 628-635. |
| [5] | Mei-ni ZUO,Yi-qing DU,Lu-ping YU,Xiang DAI,Tao XU. Correlation between metabolic syndrome and prognosis of patients with clear cell renal cell carcinoma [J]. Journal of Peking University (Health Sciences), 2022, 54(4): 636-643. |
| [6] | YANG Duo,ZHOU Xin-na,WANG Shuo,WANG Xiao-li,YUAN Yan-hua,YANG Hua-bin,GENG Hui-zhen,PENG Bing,LI Zi-bo,LI Bin,REN Jun. Assessment of lymphocytic function in vitro stimulated by specific tumor polypeptide combined with dendritic cells [J]. Journal of Peking University (Health Sciences), 2021, 53(6): 1094-1098. |
| [7] | Jing XIE,Yu-ming ZHAO,Nan-quan RAO,Xiao-tong WANG,Teng-jiao-zi FANG,Xiao-xia LI,Yue ZHAI,Jing-zhi LI,Li-hong GE,Yuan-yuan WANG. Comparative study of differentiation potential of mesenchymal stem cells derived from orofacial system into vascular endothelial cells [J]. Journal of Peking University(Health Sciences), 2019, 51(5): 900-906. |
| [8] | Fan ZHANG,Tai-qiang YAN,Wei GUO. Rasfonin inhibits proliferation and migration of osteosarcoma 143B cells [J]. Journal of Peking University(Health Sciences), 2019, 51(2): 234-238. |
| [9] | WANG Zi-cheng, CHENG Li, LV Tong-de, SU Li, LIN Jian, ZHOU Li-qun. Inflammatory priming adipose derived stem cells significantly inhibit the proliferation of peripheral blood mononuclear cells [J]. Journal of Peking University(Health Sciences), 2018, 50(4): 590-594. |
| [10] | TANG Xu, ZHAO Wei-hong, SONG Qin-qin, YIN Hua-qi, DU Yi-qing, SHENG Zheng-zuo, WANG Qiang, ZHANG Xiao-wei, LI Qing, LIU Shi-jun, XU Tao. Influence of SOX10 on the proliferation and invasion of prostate cancer cells [J]. Journal of Peking University(Health Sciences), 2018, 50(4): 602-606. |
| [11] | WANG Xiao-tong, RAO Nan-quan, FANG Teng-jiao-zi, ZHAO Yu-ming, GE Li-hong. Comparison of the properties of CD146 positive and CD146 negative subpopulations of stem cells from human exfoliated deciduous teeth [J]. Journal of Peking University(Health Sciences), 2018, 50(2): 284-292. |
| [12] | CHEN Wei, HU Fan-lei, LIU Hong-jiang, XU Li-ling, LI Ying-ni, LI Zhan-guo. Myeloid-derived suppressor cells promoted autologous B cell proliferation in rheumatoid arthritis [J]. Journal of Peking University(Health Sciences), 2017, 49(5): 819-823. |
| [13] | CAI Yi, GUO Hao, LI Han-zhong, WANG Wen-da, ZHANG Yu-shi. MicroRNA differential expression profile in tuberous sclerosis complex cell line TSC2-/- MEFs and normal cell line TSC2+/+ MEFs [J]. Journal of Peking University(Health Sciences), 2017, 49(4): 580-584. |
| [14] | SIMA Zi-han, HONG Ying-ying, LI Tie-jun△. Effects of PTCH1 mutations on the epithelial proliferation derived from keratocystic odontogenic tumour [J]. Journal of Peking University(Health Sciences), 2017, 49(3): 522-526. |
| [15] | GAO Xiang, CHEN Xiang-mei, ZHANG Ting, ZHANG Jing, CHEN Mo, GUO Zheng--yang, SHI Yan-yan, LU Feng-min, DING Shi-gang. Relationship between macrophage capping protein and gastric cancer cell’s proliferation and migration ability [J]. Journal of Peking University(Health Sciences), 2017, 49(3): 489-494. |
|
||