北京大学学报(医学版) ›› 2025, Vol. 57 ›› Issue (6): 1032-1041. doi: 10.19723/j.issn.1671-167X.2025.06.004
基于B细胞单细胞转录组测序的干燥综合征分子分型
林文灏, 谢阳, 王芳晴, 王淑盈, 刘香君, 胡凡磊*( ), 贾园*(
), 贾园*( )
)
- 北京大学人民医院风湿免疫科,北京 100044
Single-cell RNA sequencing of B cells reveals molecular typing in Sjögren syndrome
Wenhao LIN, Yang XIE, Fangqing WANG, Shuying WANG, Xiangjun LIU, Fanlei HU*( ), Yuan JIA*(
), Yuan JIA*( )
)
- Department of Rheumatology and Immunology, Peking University People ' s Hospital, Beijing 100044, China
摘要:
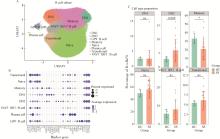
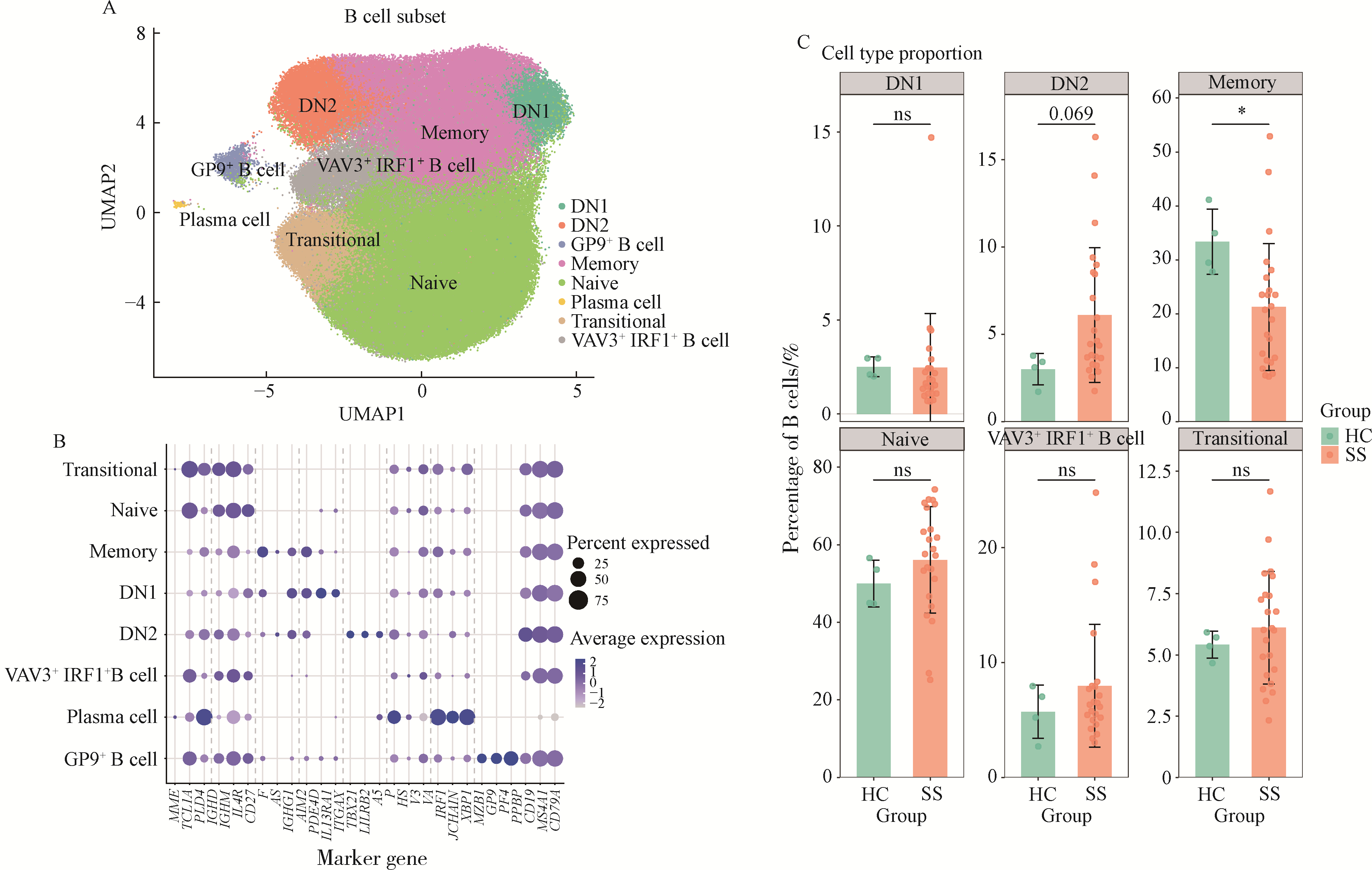
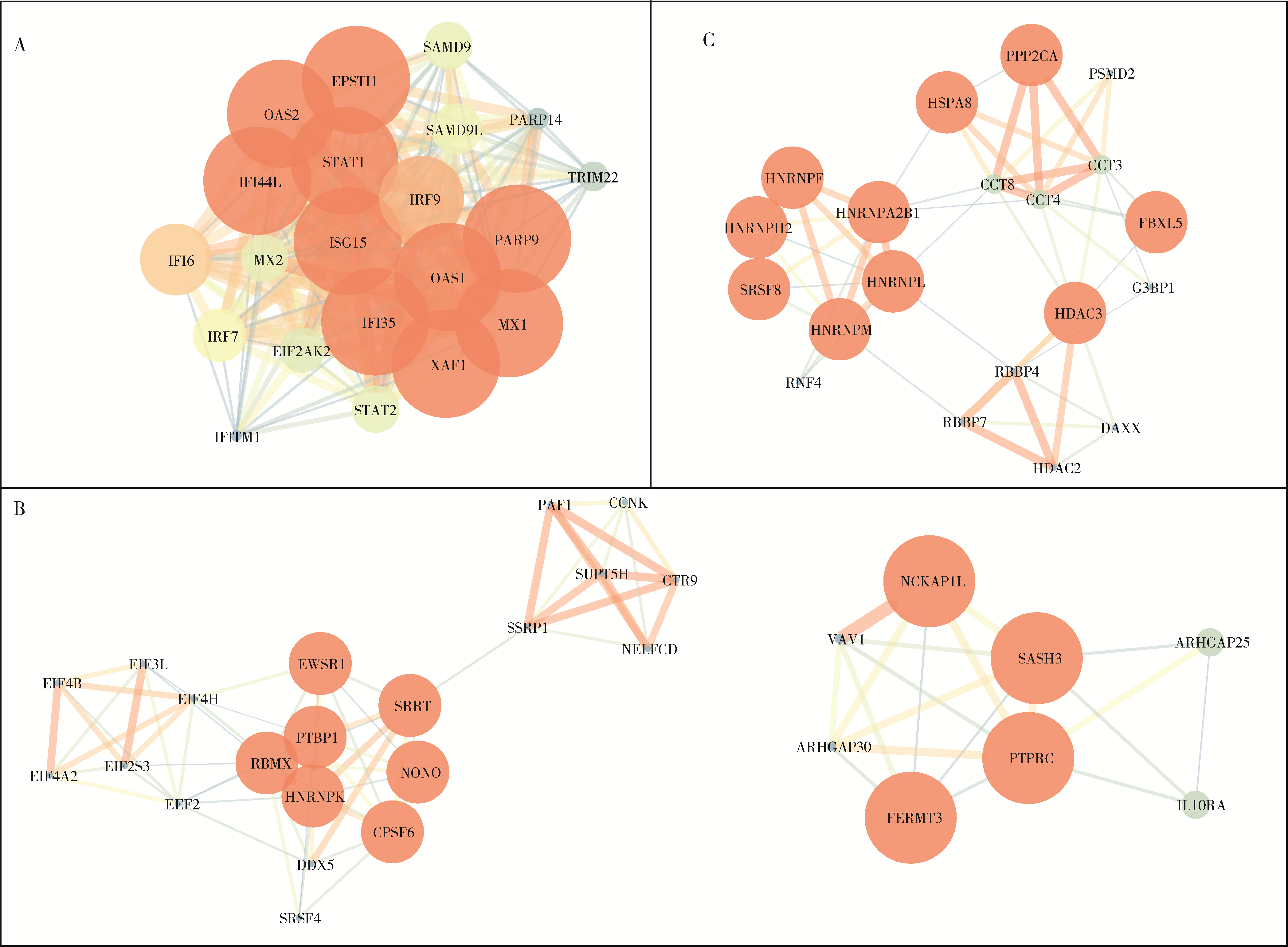
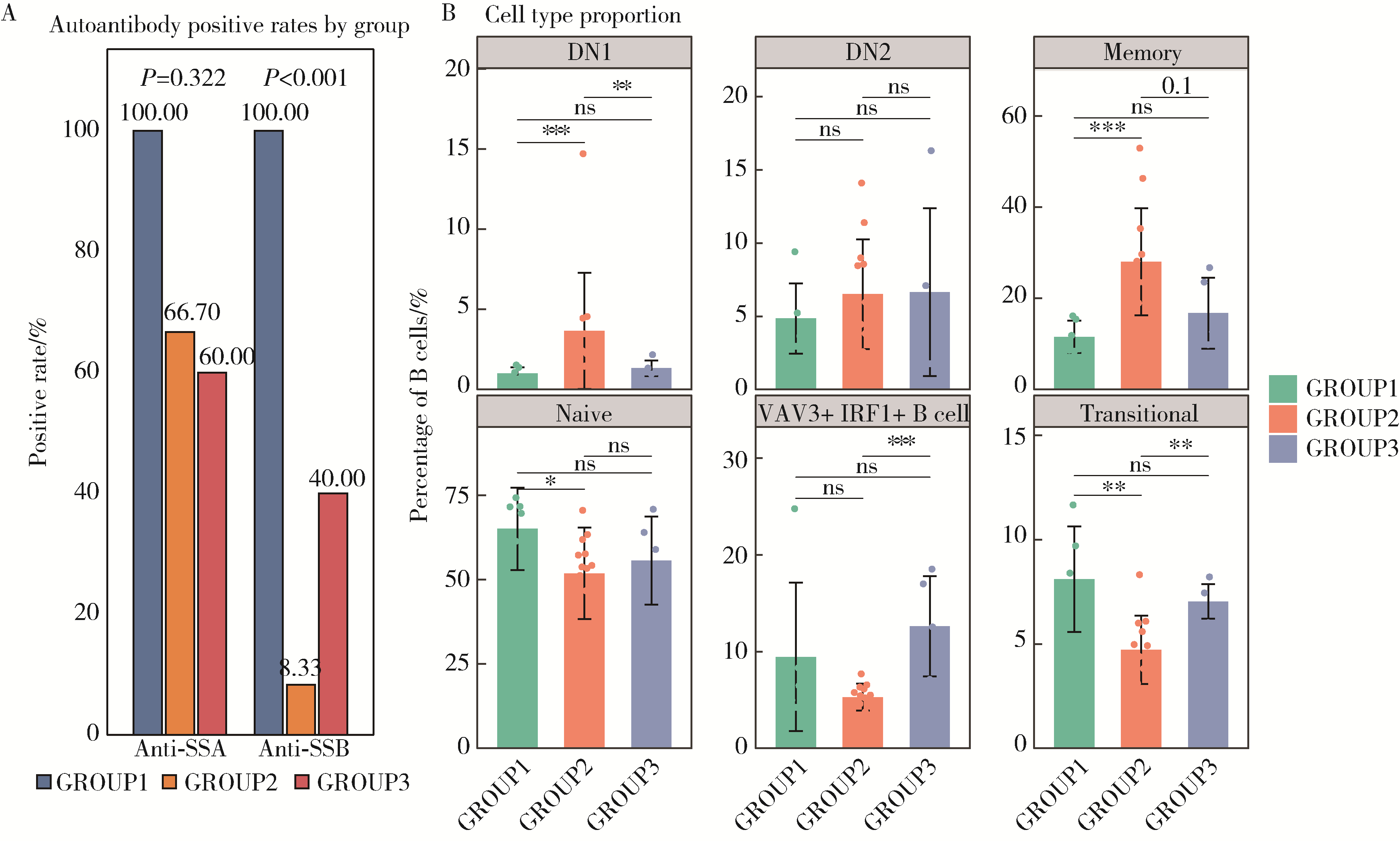
目的: 通过无监督聚类的方法分析B细胞单细胞转录组数据,将干燥综合征(Sjögren syndrome, SS)患者区分为不同亚型,建立SS的分子分型框架;同时根据不同亚型特征基因集构建蛋白质-蛋白质相互作用(protein-protein interaction,PPI)网络以探索不同亚型的核心调控机制,为精准的分子分型提供依据,并揭示SS潜在的治疗靶点,同时探索不同亚型患者的自身抗体及B细胞亚群特点。方法: 从基因表达综合数据库(Gene Expression Omnibus,GEO)获取SS患者(n=24)和健康对照(n=4)单细胞转录组测序数据,构建B细胞图谱并识别SS患者与健康对照的B细胞差异基因。通过无监督聚类将SS患者区分为不同亚型,并通过富集分析其特征基因集推测其生物学过程或通路。利用相互作用基因/蛋白质检索工具(Search Tool for the Retrieval of Interacting Genes/Proteins,STRING)数据库和Cytoscape软件构建PPI网络,推断不同亚型特征基因集的核心功能和潜在的靶点。统计并分析不同亚型患者的自身抗体情况及B细胞亚群比例。结果: 将B细胞识别为8个细胞亚群,包括过渡B细胞、初始B细胞、记忆B细胞、双阴性B细胞1型、双阴性B细胞2型、VAV3+IRF1+B细胞、GP9+B细胞和浆细胞。使用FindAllMarkers功能识别出SS患者与健康对照的B细胞差异基因792个,通过对差异基因无监督聚类分析,将SS患者分为3个亚型。通过富集分析,按照其生物学过程或通路,将3个亚型分别命名为干扰素主导亚型、B细胞活化亚型、内质网应激亚型。针对每个亚型特异的基因表达特征,构建了PPI网络,并筛选出每个亚型中的核心枢纽蛋白。每个亚型拥有不同的B细胞亚群及自身抗体特征:干扰素主导亚型有着最高的初始B细胞和过渡B细胞比例,且抗干燥综合征抗原B (Sjögren syndrome antigen B, SSB)抗体阳性率最高;B细胞活化亚型有着最高的记忆B细胞和双阴性B细胞1型比例;内质网应激亚型则有着最高的VAV3+IRF1+B细胞比例。结论: 通过无监督聚类分析SS患者与健康对照的B细胞差异表达基因,成功将SS患者分为3个分子分型,各亚型表现出特征性的自身抗体谱和优势B细胞亚群,此分子分型框架揭示了SS的B细胞特征,为探索潜在治疗靶点和指导精准治疗提供了新的依据。
中图分类号:
- R593.2
| 1 |
中华医学会风湿病学分会. 干燥综合征诊断及治疗指南[J]. 中华风湿病学杂志, 2010, 14(11): 766- 768.
|
| 2 |
doi: 10.1038/nrrheum.2018.1 |
| 3 |
doi: 10.1016/j.immuni.2009.11.009 |
| 4 |
doi: 10.1002/art.42683 |
| 5 |
doi: 10.1186/s13073-023-01237-9 |
| 6 |
doi: 10.3389/fmolb.2023.1202371 |
| 7 |
doi: 10.1002/advs.202417593 |
| 8 |
doi: 10.1038/s41592-019-0619-0 |
| 9 |
|
| 10 |
doi: 10.1038/nbt.4096 |
| 11 |
doi: 10.1093/nar/gkac1000 |
| 12 |
doi: 10.1101/gr.1239303 |
| 13 |
doi: 10.1186/1471-2105-4-2 |
| 14 |
doi: 10.1016/S2665-9913(19)30042-6 |
| 15 |
doi: 10.1038/s41467-021-23472-7 |
| 16 |
doi: 10.1136/annrheumdis-2021-221604 |
| 17 |
doi: 10.1007/s10067-025-07409-9 |
| 18 |
doi: 10.3390/ijms24098385 |
| 19 |
doi: 10.1038/nri856 |
| 20 |
doi: 10.1186/s42358-024-00430-7 |
| 21 |
doi: 10.1016/j.canlet.2024.217045 |
| 22 |
doi: 10.2147/JIR.S413581 |
| 23 |
doi: 10.1038/374470a0 |
| 24 |
doi: 10.2147/JIR.S440263 |
| 25 |
doi: 10.1371/journal.pbio.3000252 |
| 26 |
杜晶晶, 董兴红, 高洁, 等. 原发性干燥综合征患者抗SSB与其他实验室参数相关性研究[J]. 检验医学与临床, 2023, 20(16): 2395- 2399.
|
| 27 |
doi: 10.1080/07853890.2022.2031272 |
| 28 |
doi: 10.1016/j.jtauto.2024.100252 |
| 29 |
doi: 10.1038/s41467-022-30651-7 |
| [1] | 刘源, 石桂秀. 干燥综合征到干燥病的命名变迁[J]. 北京大学学报(医学版), 2025, 57(6): 1015-1017. |
| [2] | 向钊, 杨莉, 杨静. 非靶向代谢组学揭示原发性干燥综合征血小板减少患者血清差异代谢物及代谢通路[J]. 北京大学学报(医学版), 2025, 57(6): 1042-1050. |
| [3] | 赵亚云, 倪梦凡, 李雪, 王蓓, 程功, 何菁, 金月波. 利妥昔单抗治疗原发性干燥综合征肾损害的临床疗效和安全性[J]. 北京大学学报(医学版), 2025, 57(6): 1051-1060. |
| [4] | 朱丽秀, 陈仁利, 周素娟, 林烨, 汤一榕, 叶桢. 水通道蛋白5对干燥综合征大鼠TLR4/MyD88/NF-κB信号的影响[J]. 北京大学学报(医学版), 2025, 57(5): 875-883. |
| [5] | 宁圆, 张晓盈, 李雪, 李原, 何菁, 金月波. 干燥综合征并发乳腺淋巴瘤1例[J]. 北京大学学报(医学版), 2025, 57(4): 808-811. |
| [6] | 杨玉淑, 齐晅, 丁萌, 王炜, 郭惠芳, 高丽霞. 抗唾液腺蛋白1抗体联合抗腮腺分泌蛋白抗体对干燥综合征的诊断价值[J]. 北京大学学报(医学版), 2024, 56(5): 845-852. |
| [7] | 蔡祥,王仁东,王世佳,任梓齐,于秋红,李冬果. 胶质母细胞瘤恶性进展中不同细胞亚群的动态轨迹和细胞通讯网络[J]. 北京大学学报(医学版), 2024, 56(2): 199-206. |
| [8] | 韩艺钧,李常虹,陈秀英,赵金霞. 抗SSB抗体阳性和阴性的原发性干燥综合征患者临床及免疫学特征的比较[J]. 北京大学学报(医学版), 2023, 55(6): 1000-1006. |
| [9] | 李建斌,吕梦娜,池强,彭一琳,刘鹏程,吴锐. 干燥综合征患者发生重症新型冠状病毒肺炎的早期预测[J]. 北京大学学报(医学版), 2023, 55(6): 1007-1012. |
| [10] | 孟彦宏,陈怡帆,周培茹. CENP-B抗体阳性的原发性干燥综合征患者的临床和免疫学特征[J]. 北京大学学报(医学版), 2023, 55(6): 1088-1096. |
| [11] | 吴洁,张雯,梁舒,秦艺璐,范文强. 妊娠期原发性干燥综合征合并视神经脊髓炎谱系疾病危重症1例[J]. 北京大学学报(医学版), 2023, 55(6): 1118-1124. |
| [12] | 王丽芳,石连杰,宁武,高乃姝,王宽婷. 干燥综合征合并冷凝集素病1例[J]. 北京大学学报(医学版), 2023, 55(6): 1130-1134. |
| [13] | 邢海霞,王琳,乔迪,刘畅,潘洁. 干燥综合征口腔疾病的治疗特点[J]. 北京大学学报(医学版), 2023, 55(5): 929-933. |
| [14] | 蔡文心,李仕成,刘一鸣,梁如玉,李静,郭建萍,胡凡磊,孙晓麟,李春,刘栩,叶华,邓立宗,李茹,栗占国. 类风湿关节炎临床分层及其特征的横断面研究[J]. 北京大学学报(医学版), 2022, 54(6): 1068-1073. |
| [15] | 刘杨,程昉,王艳玲,艾香艳,朱振航,赵福涛. 唾液腺超声对干燥综合征的诊断价值[J]. 北京大学学报(医学版), 2022, 54(6): 1123-1127. |
|
||